Performance Assessment of Variant Calling Pipelines using Human Whole Exome Sequencing and Simulated data | bioRxiv
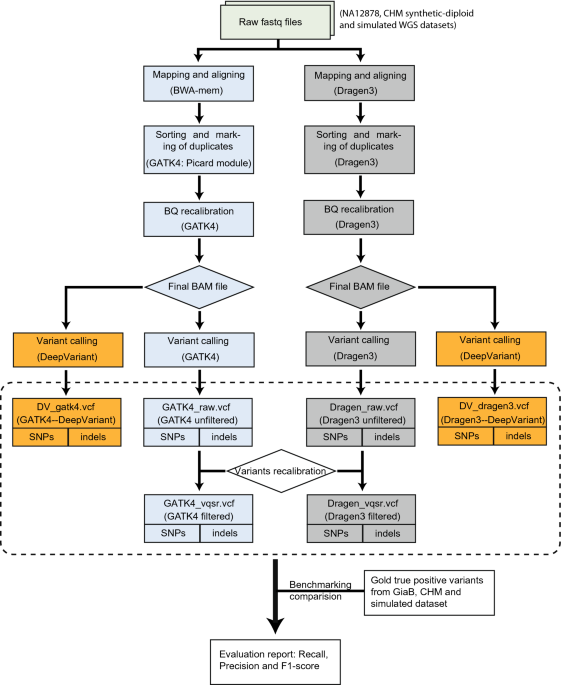
Accuracy and efficiency of germline variant calling pipelines for human genome data | Scientific Reports
Performance comparison of seven variant calling tools. The performance... | Download Scientific Diagram

Calling Variants in the Clinic: Informed Variant Calling Decisions Based on Biological, Clinical, and Laboratory Variables - Computational and Structural Biotechnology Journal
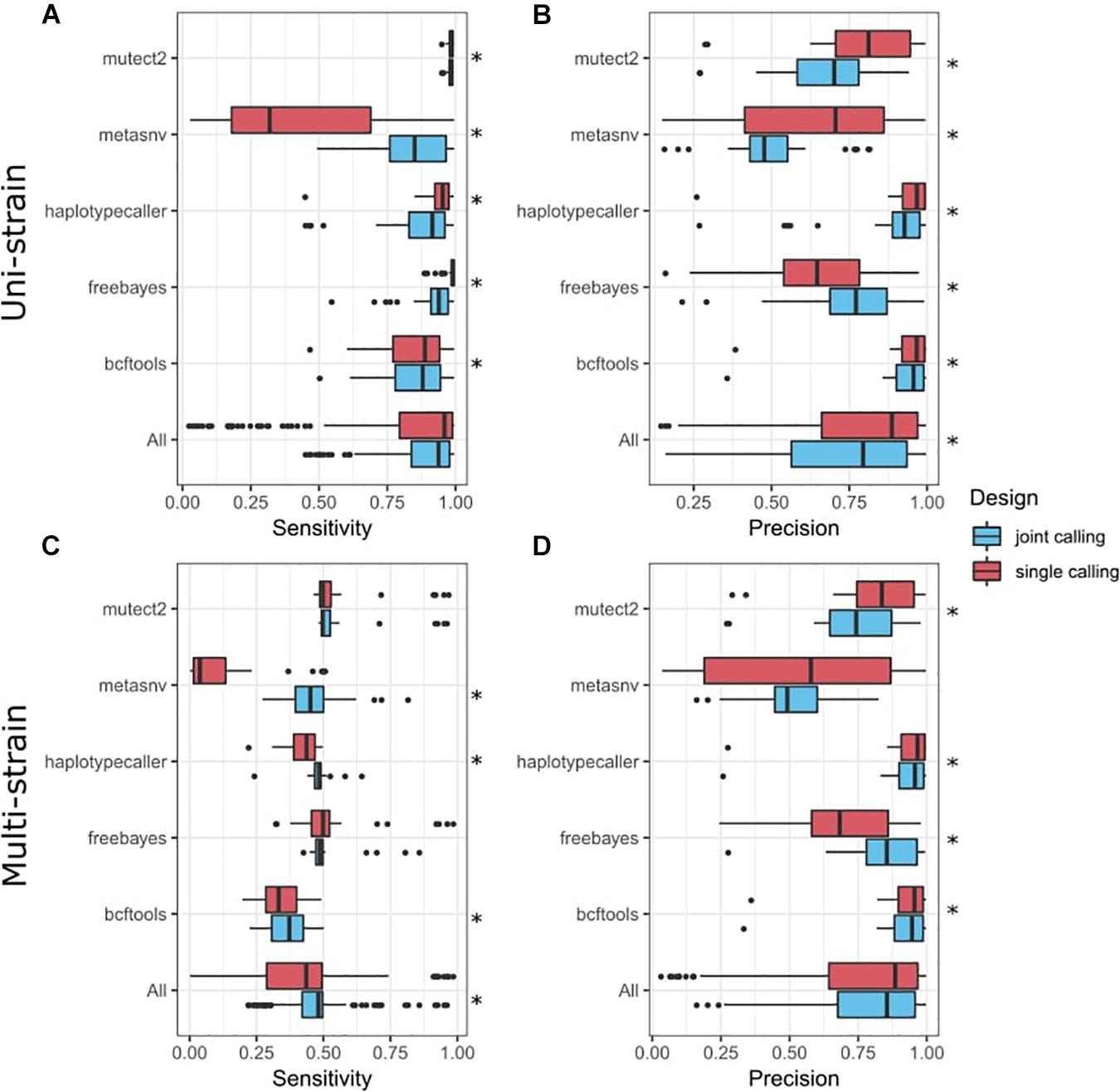
Frontiers | A Benchmark of Genetic Variant Calling Pipelines Using Metagenomic Short-Read Sequencing

Best practices for benchmarking germline small-variant calls in human genomes | Nature Biotechnology
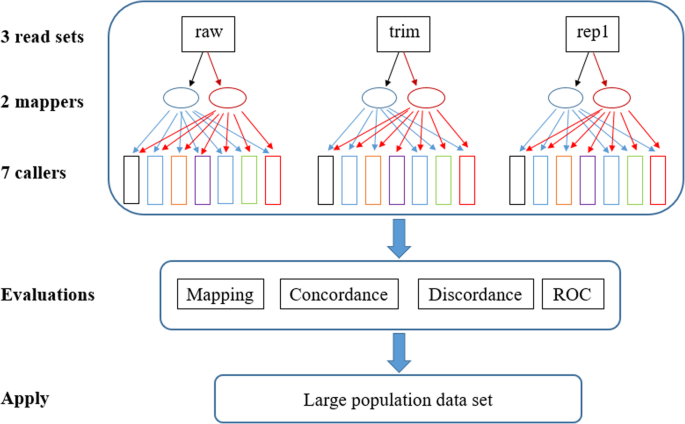
Evaluation of variant calling tools for large plant genome re-sequencing | BMC Bioinformatics | Full Text

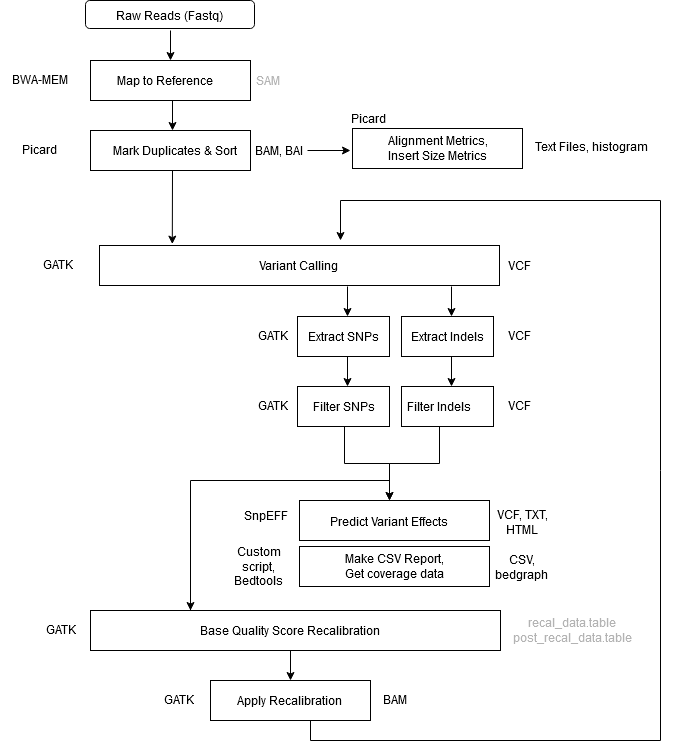
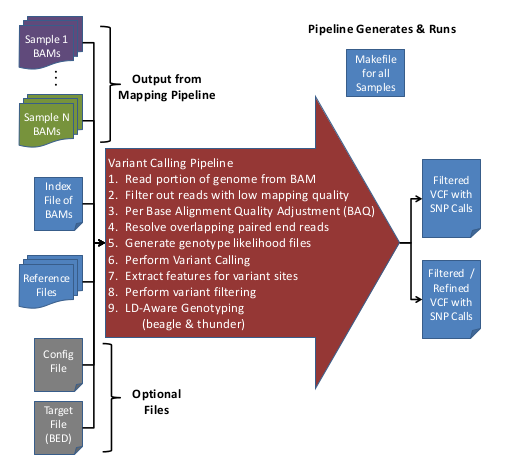
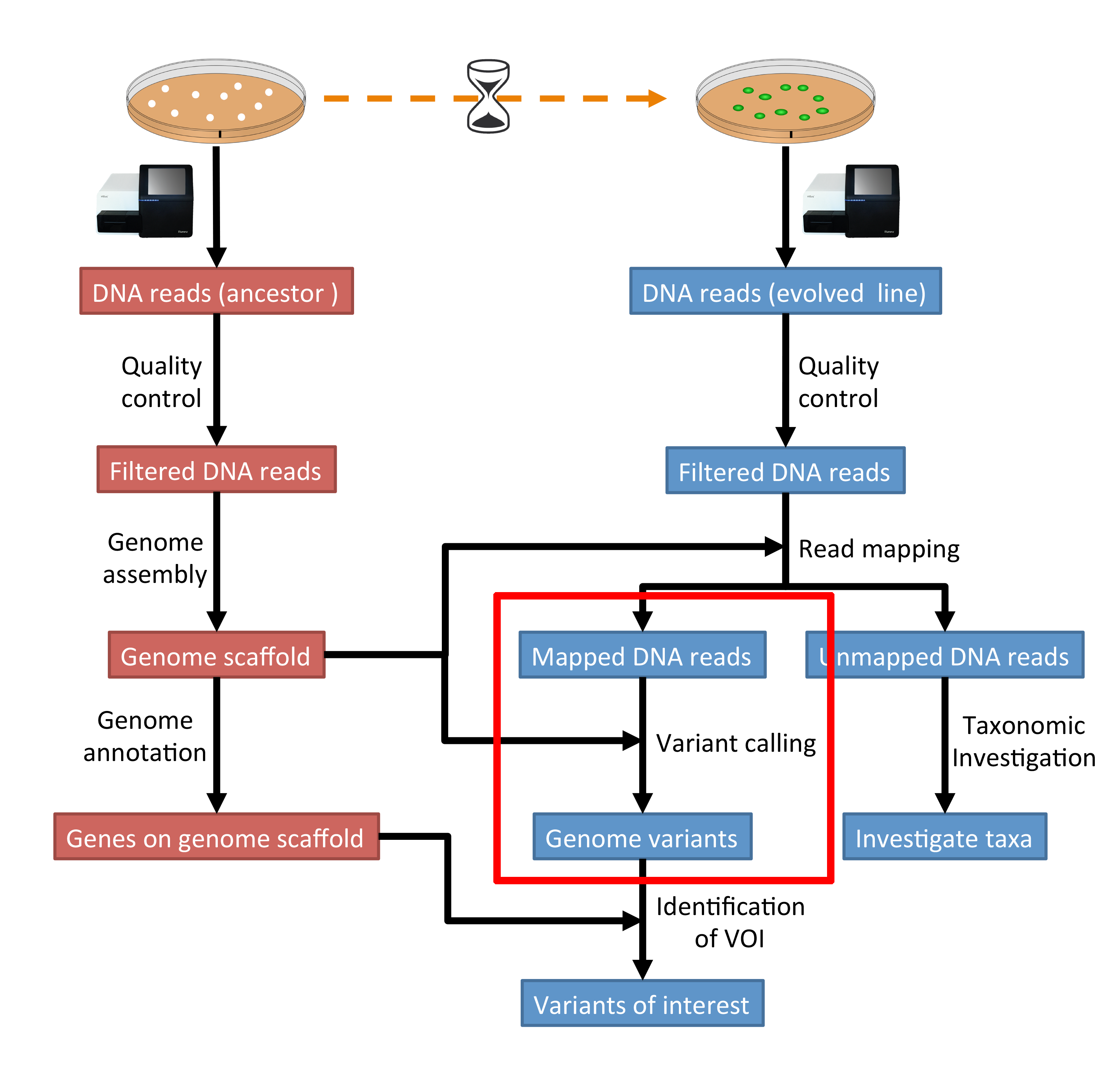
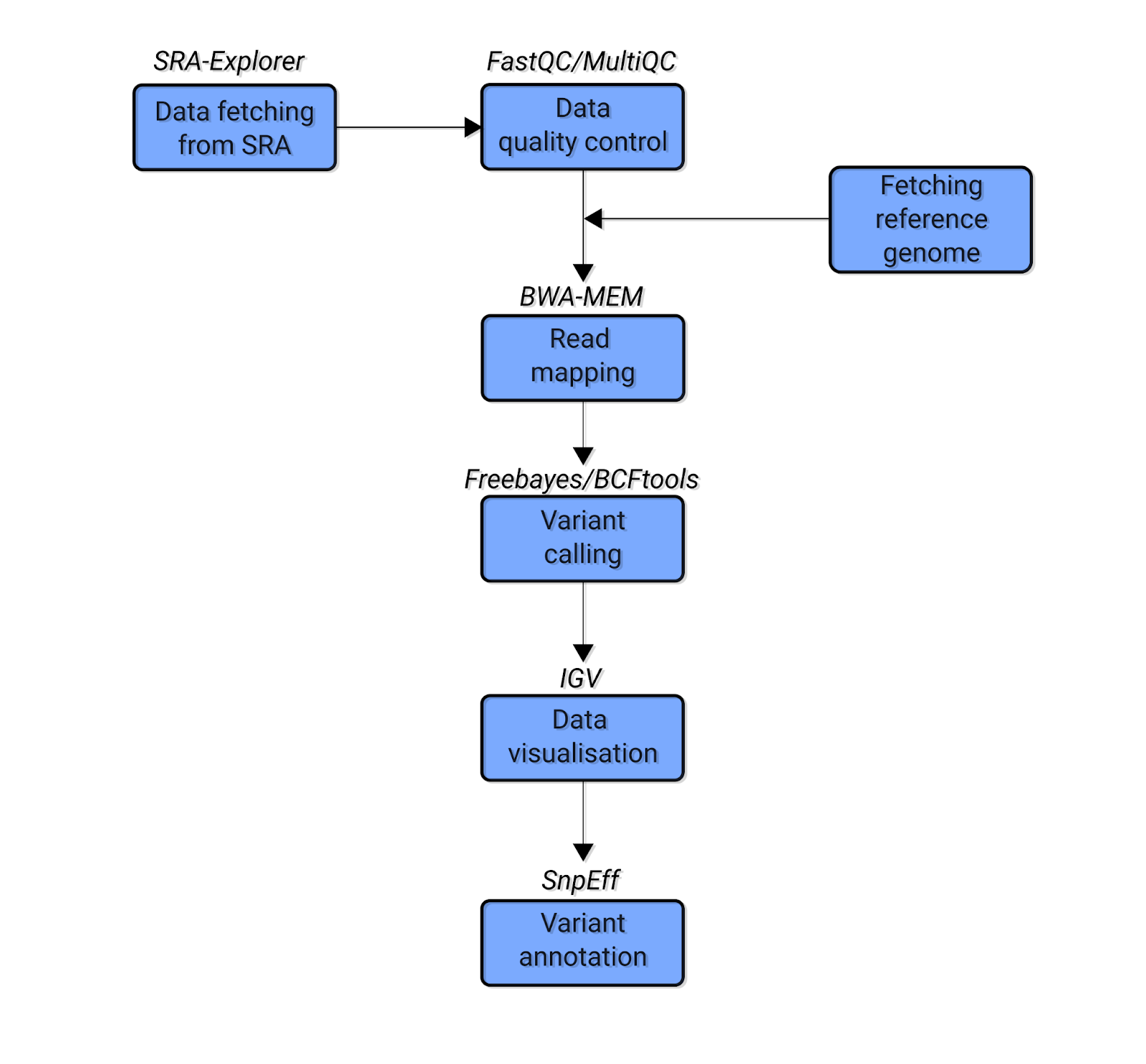
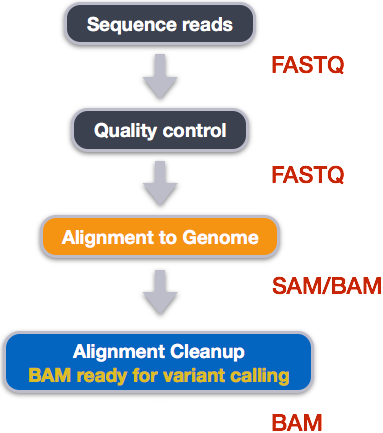
Variant calling pipeline. Schematic representation of the bioinformatic... | Download Scientific Diagram